Multi-omics analysis of niche specificity provides new insights into ecological adaptation in bacteria
The ISME Journal, a series under Nature Publishing Group, published an article entitled Multi-omics analysis of niche specificity provides new insights into ecological adaptation in bacteria in October, 2016. The article reported the recent research progress on new mechanisms of bacterial adaptation in micro-environment accomplished jointly by groups headed by Prof. ZHOU Xueping, IPP-CAAS, and Associated Prof. LI Bin in College of Agriculture and Biotechnology, Zhejiang University, respectively.
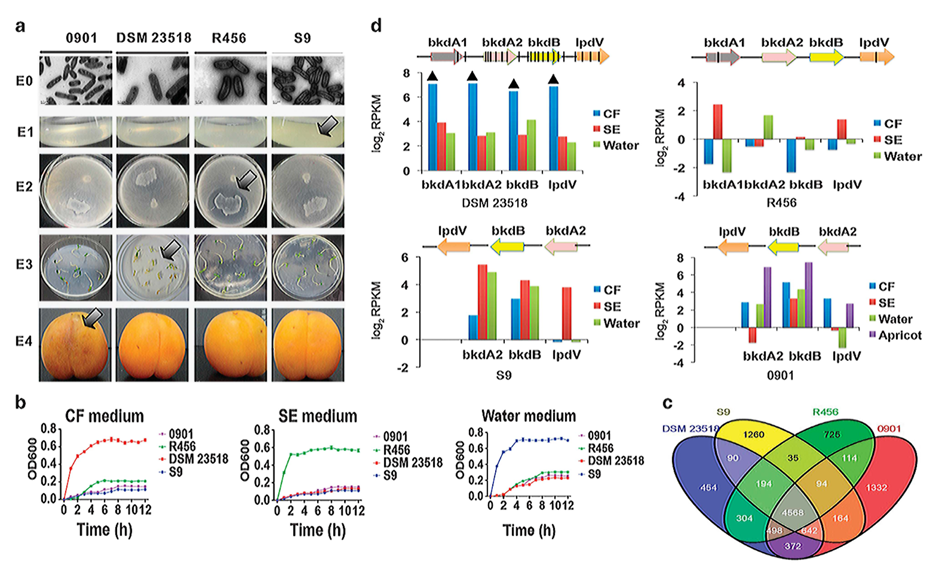
It has been a hot spot for evolutional bacteriology research that different closely related bacterial strains can adapt to entirely different niche. Started from this, the two teams used strains 0901, S9, R456 and LMG24067 in Burkholderias seminalis as materials that respectively played roles of plant pathogen, saprophyte, biologic control agent of rice diseases and human pathogen. The scientists not only discovered the function of locus specificity in the forming of the ecological roles of these strains, but also analyzed the molecular mechanism of these closely related strains adapting to different niche by using phenomic, genomic, transcriptomic and methylomic methods.
The results indicated that each strain is separated from the others by increased fitness in medium simulating its original niche corresponding to the difference between strains in metabolic capacities. Strain-specific metabolism and niche survival was generally linked with genomic variants and niche-dependent differential expression of the corresponding genes. For example, large number of tandem genes that were plant pathogen specific was discovered only in genome of the plant pathogen. No homologous genes were discovered in the other three strains. This showed obtained gene islands via horizontal gene transfer. The analysis of expression revealed a group of gene islands that had been reported as closely related to plant pathogenicity had statistically high expression while the pathogen infected plant. In particular, the importance of iron, trehalose and D-arabitol utilization was highlighted by the involvement of DNA-methylation and in niche-adapted regulation of the corresponding operons based on the integrated analysis of their multi-omics data. Overall, the results provided insights of niche-specific adaptation in bacteria.
The link to the article: http://www.nature.com/ismej/journal/vaop/ncurrent/abs/ismej2015251a.html
-
 China-Laos Training Workshop on Integrated Management of Destructive Crop Pests and Diseases Successfully held in Laos
China-Laos Training Workshop on Integrated Management of Destructive Crop Pests and Diseases Successfully held in Laos -
 New Plant Protection: New challenge and new opportunity for plant protection
New Plant Protection: New challenge and new opportunity for plant protection -
 International and SELAMAT Conference on Pesticide Residue and Mycotoxin Contamination Held in Beijing
International and SELAMAT Conference on Pesticide Residue and Mycotoxin Contamination Held in Beijing -
 CAAS President Meets Chairman of ASEAN FAW Taskforce
CAAS President Meets Chairman of ASEAN FAW Taskforce
